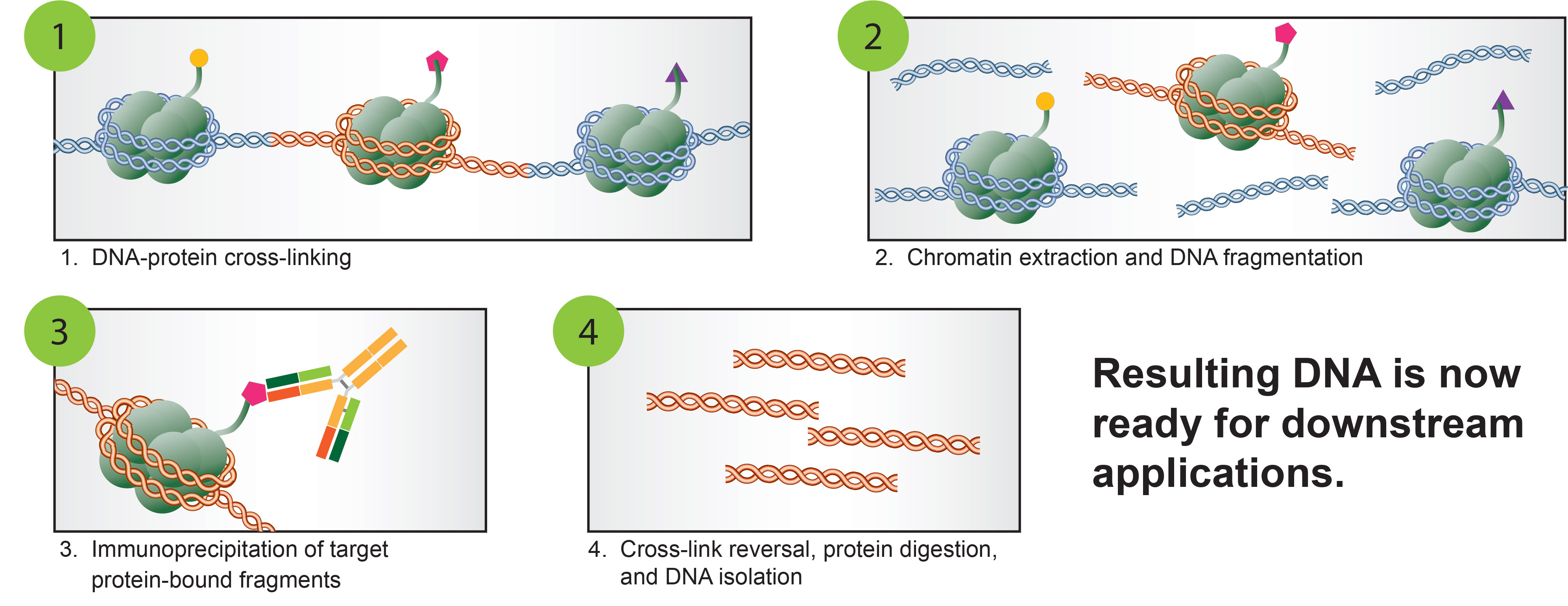
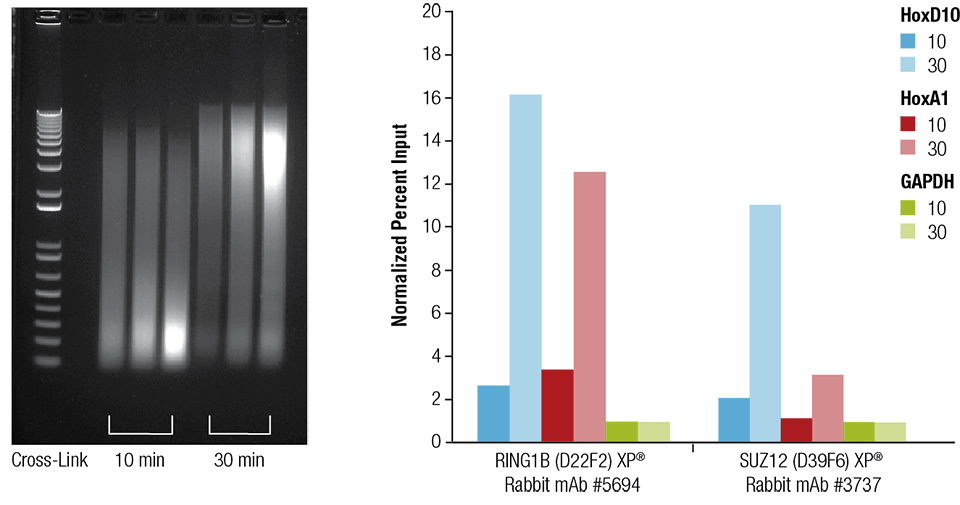
An optimized protocol for rapid, sensitive and robust on-bead ChIP-seq from primary cells - ScienceDirect
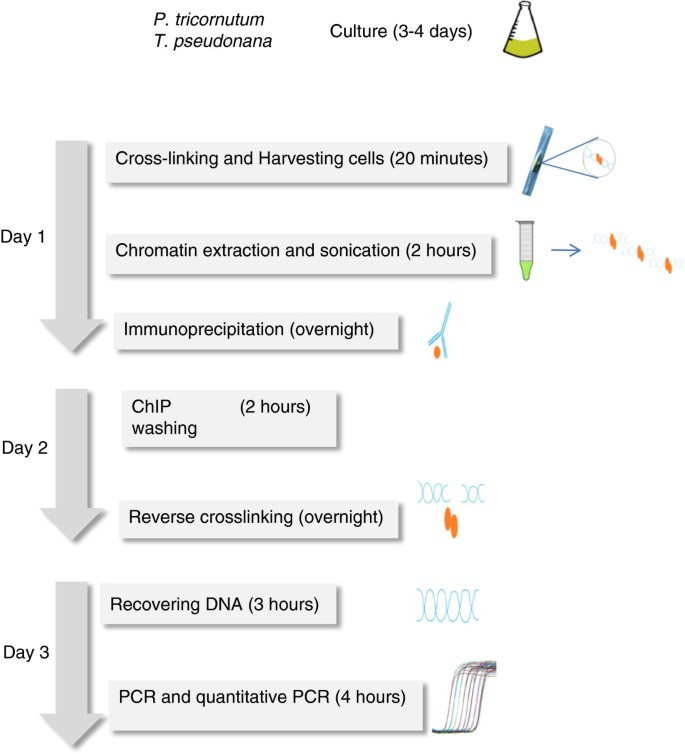
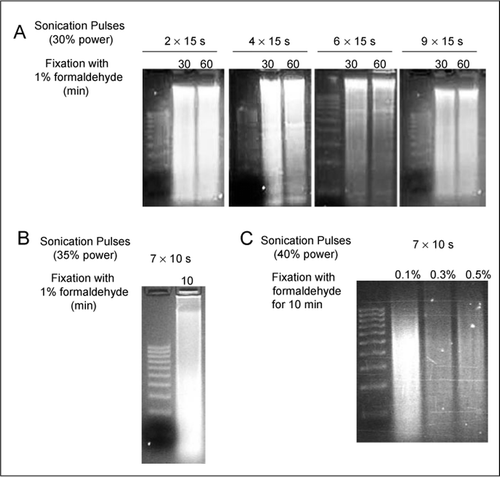
Protocol: Chromatin immunoprecipitation (ChIP) methodology to investigate histone modifications in two model diatom species | Plant Methods | Full Text
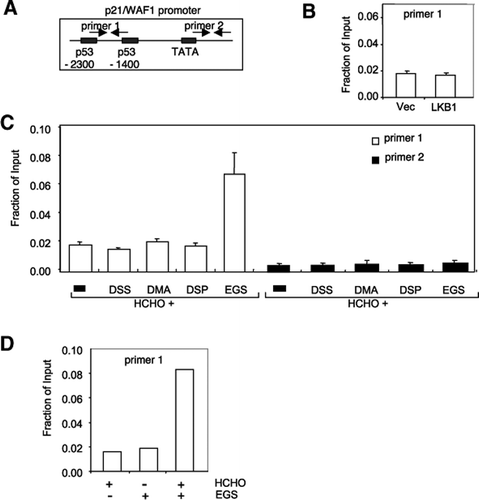
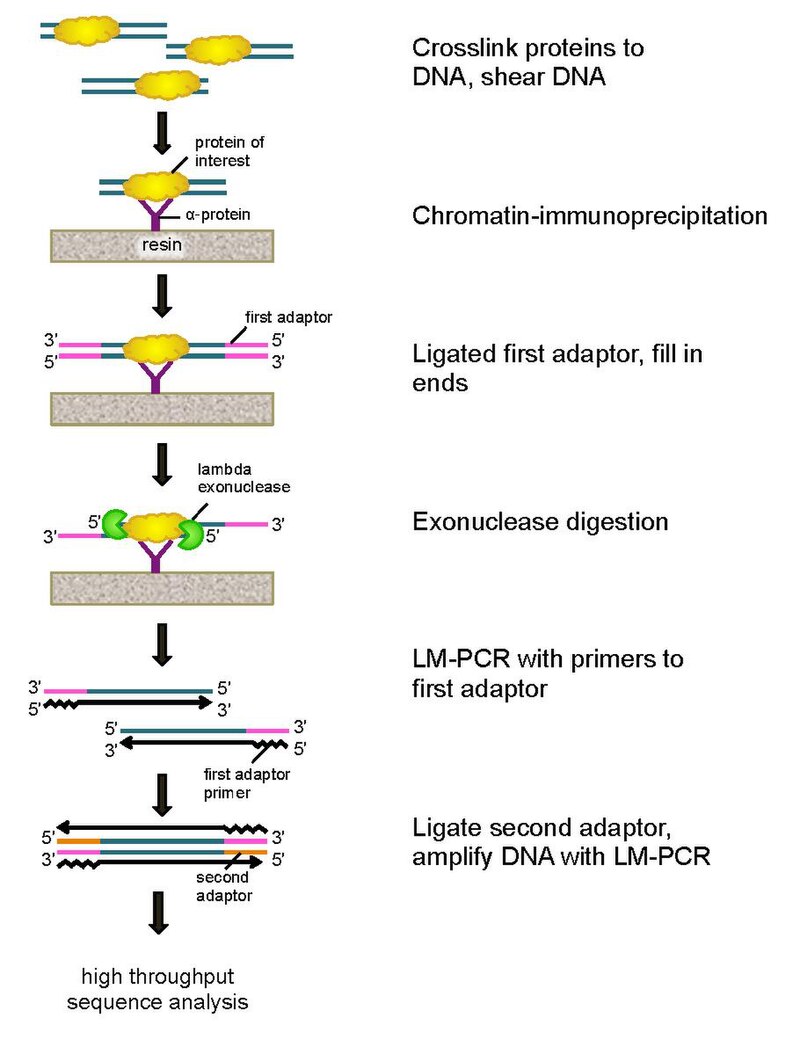
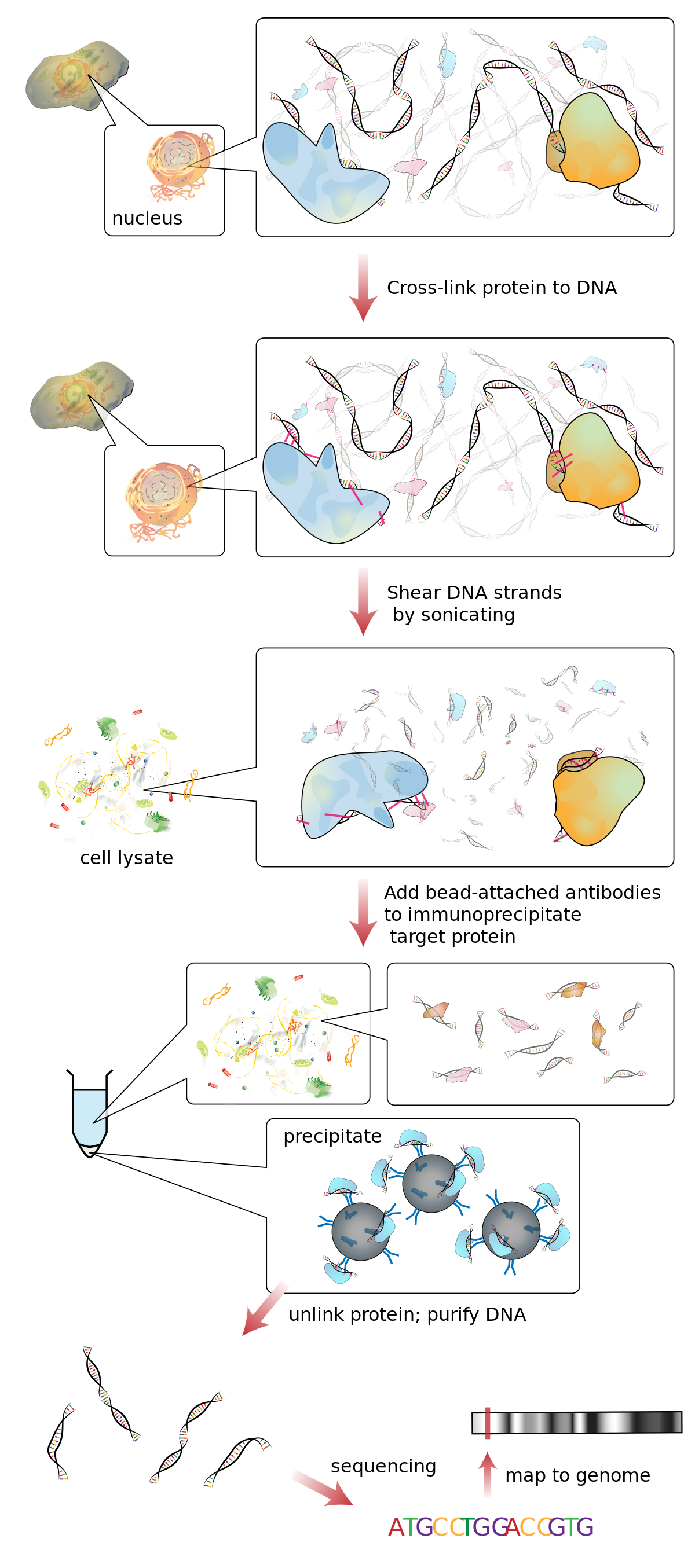
In vivo dual cross-linking for identification of indirect DNA-associated proteins by chromatin immunoprecipitation | BioTechniques
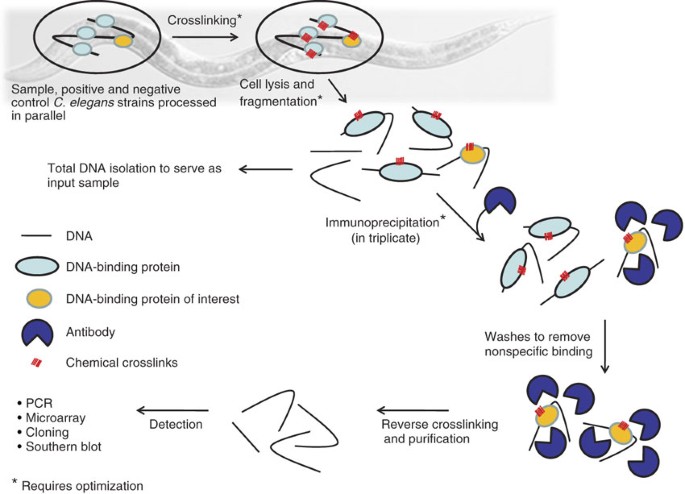
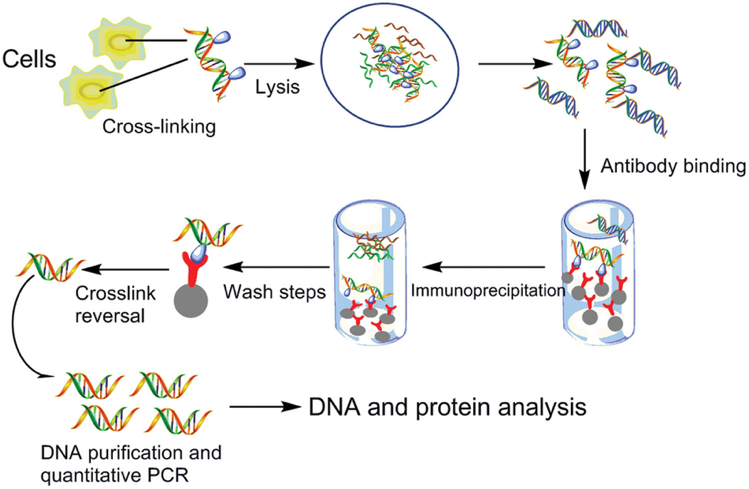
Chromatin immunoprecipitation (ChIP) coupled to detection by quantitative real-time PCR to study transcription factor binding to DNA in Caenorhabditis elegans | Nature Protocols

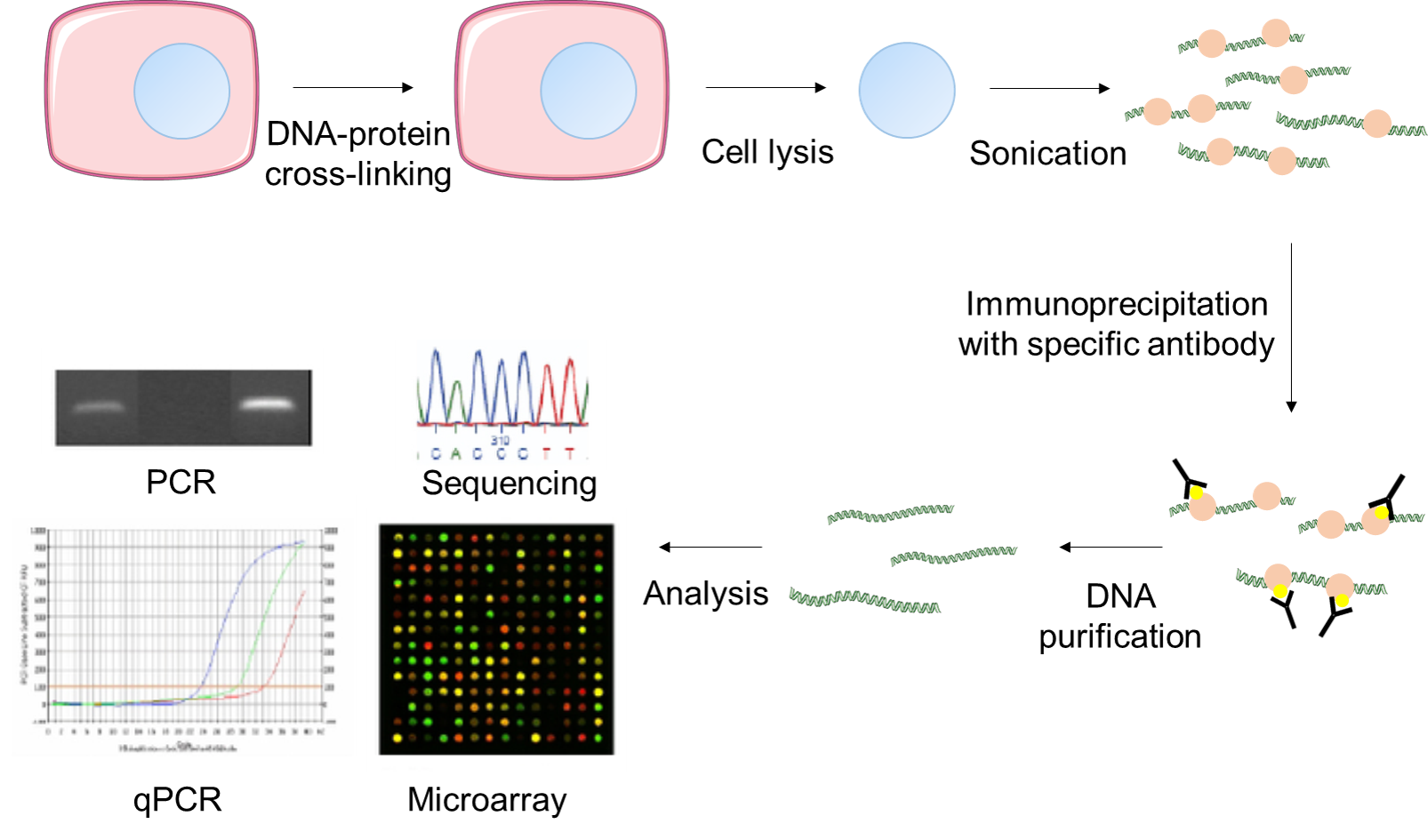
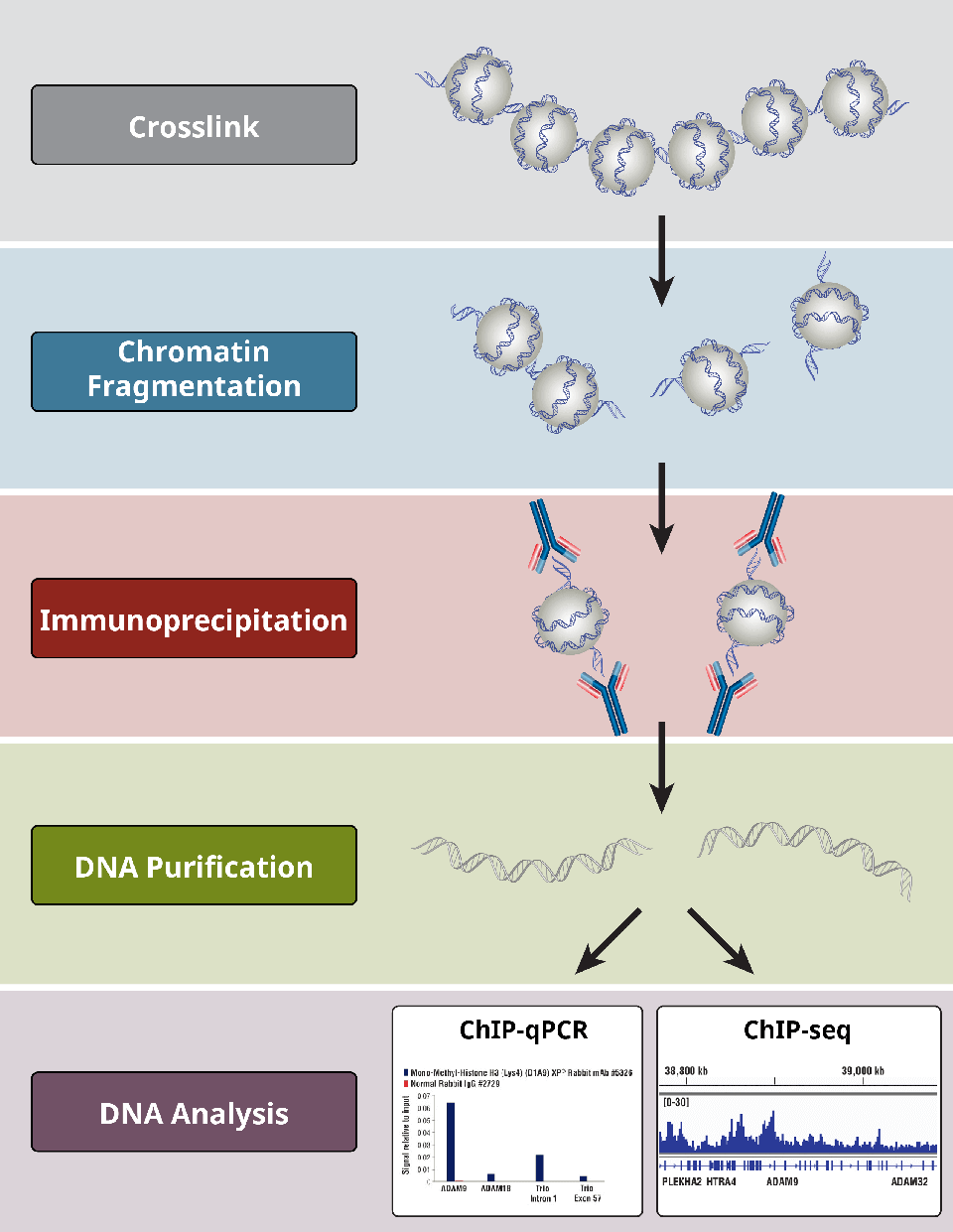
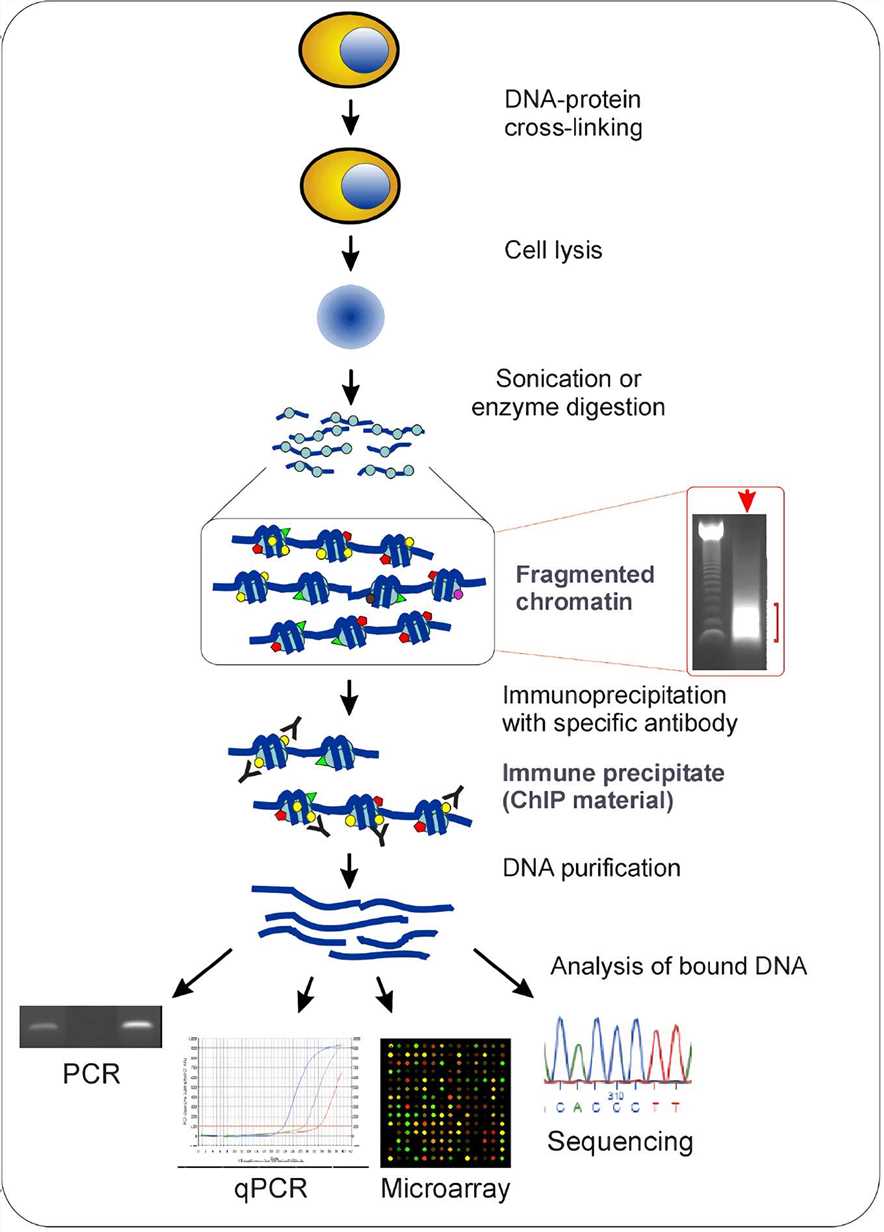
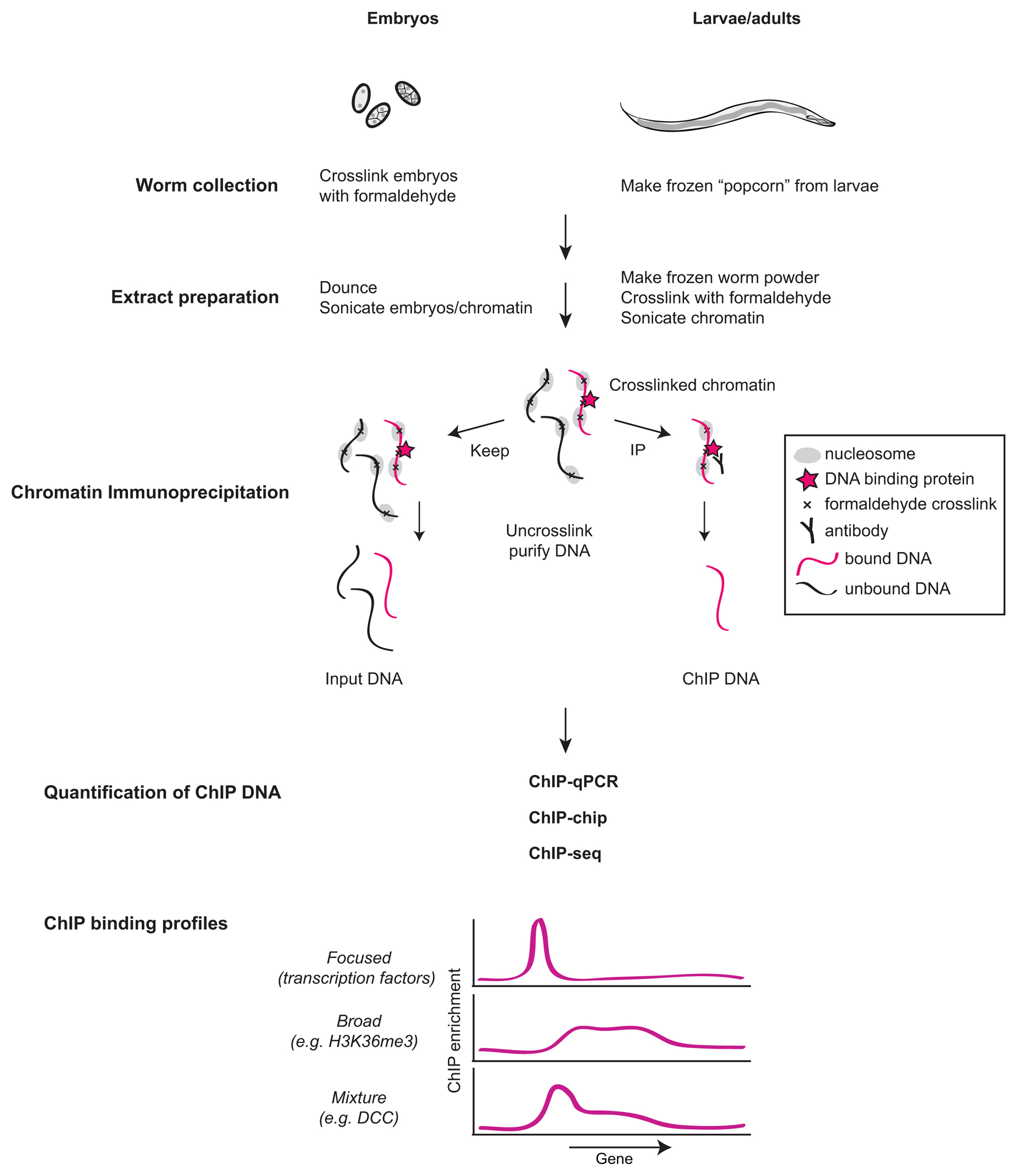
Outline of the ChIP procedure. After cross-linking with formaldehyde,... | Download Scientific Diagram

Outline of the ChIP procedure. After cross-linking with formaldehyde,... | Download Scientific Diagram